Our Research

We live in a cyclic environment; days and nights are a common thing to us. We are used to sunrises and sunsets and yet, we normally forget that such constant rhythmicity has sculpted organismal biology since the dawn of times. Biological systems have thus evolved sophisticated molecular devices that allow them to anticipate such daily changes.
Circadian (circa diem) clocks are accurate pacemakers (with ~24 h period length) that allow temporal compartmentalization of a large range of biological process within the cell and also at the organismal level. Interestingly, although the molecular components of circadian clocks differ across phyla, they share the same basic design or blueprints. Therefore, in organisms as diverse as humans, insects and fungi, it is possible to recognize a transcriptional-translational negative feedback loop at the heart of the timing mechanism, with kinases playing a key role in setting up the speed of the clock.
Circadian (circa diem) clocks are accurate pacemakers (with ~24 h period length) that allow temporal compartmentalization of a large range of biological process within the cell and also at the organismal level. Interestingly, although the molecular components of circadian clocks differ across phyla, they share the same basic design or blueprints. Therefore, in organisms as diverse as humans, insects and fungi, it is possible to recognize a transcriptional-translational negative feedback loop at the heart of the timing mechanism, with kinases playing a key role in setting up the speed of the clock.
The Larrondo Lab is interested in different aspects of gene expression. In particular, we study the molecular mechanisms underlying biological oscillators, assessing the impact that circadian clocks have on physiology and in host-pathogen interactions.
Currently, we study various fungal systems to try to understand some of these processes: the filamentous fungus Neurospora crassa, the phytopathogen Botrytis cinerea, the biocontroller Trichoderma atroviride and the yeast Saccharomyces cerevisiae.
Lately, through both optogenetics and synthetic biology approaches, we are additionally exploring the design of new oscillatory circuits capable of starting and sustaining circadian rhythms, while also developing new tools to implement complex circuits to reprogram gene expression.
Lately, through both optogenetics and synthetic biology approaches, we are additionally exploring the design of new oscillatory circuits capable of starting and sustaining circadian rhythms, while also developing new tools to implement complex circuits to reprogram gene expression.
The Neurospora circadian clock: an overview
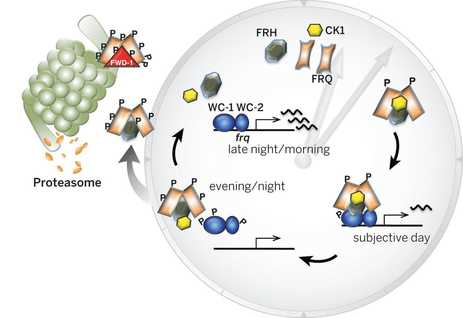
The Neurospora circadian oscillator is based on a transcriptional/translational negative feedback loop (TTFL) that relies on the negative element FREQUENCY (FRQ). frq expression is under the control of the positive element, the WCC transcriptional complex, which is composed by the GATA type transcription factors (TFs) White Collar 1 (WC-1, which is also a blue light receptor) and White Collar 2 (WC-2). Under constant darkness (DD) WCC binds to the clock-box (c-box), a cis-element within the frq promoter that is both necessary and sufficient for frq rhythmic expression. In the presence of light, the stoichiometry of the WCC changes, which alters its DNA recognition patterns, which results in the WCC binding a different cis-sequence in the frq promoter -the proximal light regulatory element, pLRE-, increasing frq expression and allowing entrainment of the clock to environmental light (summarized in Montenegro-Montero et al., 2015).
The Neurospora circadian oscillator is based on a transcriptional/translational negative feedback loop (TTFL) that relies on the negative element FREQUENCY (FRQ). frq expression is under the control of the positive element, the WCC transcriptional complex, which is composed by the GATA type transcription factors (TFs) White Collar 1 (WC-1, which is also a blue light receptor) and White Collar 2 (WC-2). Under constant darkness (DD) WCC binds to the clock-box (c-box), a cis-element within the frq promoter that is both necessary and sufficient for frq rhythmic expression. In the presence of light, the stoichiometry of the WCC changes, which alters its DNA recognition patterns, which results in the WCC binding a different cis-sequence in the frq promoter -the proximal light regulatory element, pLRE-, increasing frq expression and allowing entrainment of the clock to environmental light (summarized in Montenegro-Montero et al., 2015).
|
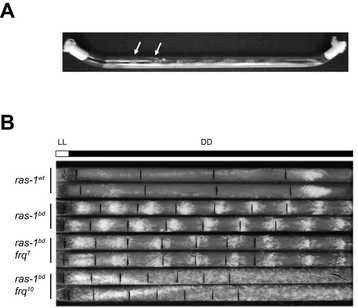
When FRQ is produced, it associates with FRH, a RNA-helicase, and this complex blocks the activity of the WCC, thus creating a negative feedback loop (that’s why FRQ is called the negative element of the clock). Shortly after its synthesis, FRQ is progressively phosphorylated up until the point in which its ability to repress the WCC is lost and FRQ eventually becomes degraded via the proteosome. Each cycle of this TTFL takes about 22.5 hours, which can be observed as daily oscillations of frq and FRQ levels and as rhythmic gene expression of 10-40% of its transcriptome, impacting, for instance, metabolism and leading to overt rhythms in conidiation. These rhythms can easily be tracked through a phenotypic assay based on “race tubes”, which allows visualization of daily rhythms in conidia production (visible as “bands” of spores, see figure on the right).
Additional information on the Neurospora circadian clock, a field that encompasses the work of several labs for over 25 years, can be found in Montenegro-Montero et al. (2015), Larrondo & Canessa (2019) and references therein. |
From Montenegro-Montero A & Larrondo LF (2013). The Neurospora Circadian System: From Genes to Proteins and Back in Less Than 24 hours, in Neurospora: Genomics and Molecular Biology.
|
Current Projects
|
I. Neurospora clock mechanisms
An important advancement in the past years has been the use of luciferase-based real-time reporters to monitor circadian gene expression in Neurospora (Gooch et al., 2008). This, together with the ease of modifying loci at will through allele replacement strategies (Larrondo et al., 2009), has allowed to further track FRQ dynamics with high spatial and temporal resolution (Larrondo et al., 2012). With such tools, we have been able to redefine some long-standing paradigms regarding period determination and FRQ stability, showing that FRQ posttranslational modifications (the quality of FRQ) rather than its quantity and degradation is crucial for determining period length (Larrondo et al., 2015; Montenegro-Montero et al., 2015). Current efforts are focused on understanding the intrincate mechanistic connections between FRQ dynamics, CK1 activity and period determination. Indeed, our results highlight the relevance of CK1 on the core-clock, while also providing solid evidence that other kinases, such as PKA, have minimal impact on circadian parameters (Olivares-Yanez et al. 2023). |
From Larondo et al., (2015). Circadian rhythms. Decoupling circadian clock protein turnover from circadian period determination.
Science. 347(6221):1257277 |
|
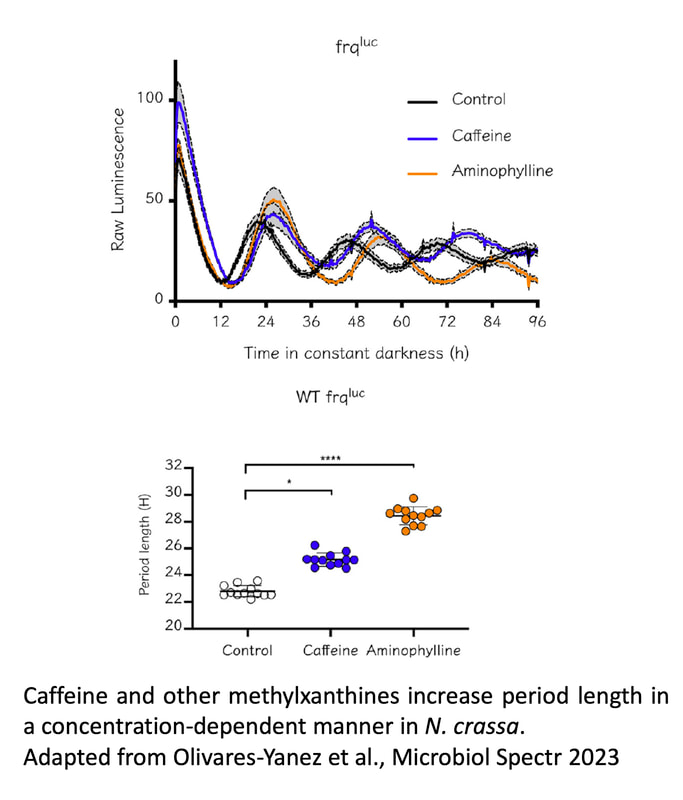
We are also interested in other aspects of circadian regulation in Neurospora, such as the cross-talk between metabolism and clock regulation and its effect on cellulose degradation (Diaz & Larrondo 2020), or mechanisms underlying metabolic compensation (Olivares-Yañez et al., 2016). We have also studied the role of second messengers (such as cAMP) and the effect of several drugs on clock dynamics, such as caffeine and other methylxantines (Olivares-Yanez et al. 2023), together with actively examining the role of diverse transcription factors in regulating not only core-clock function, but also circadian output pathways which are also subjected to postranscriptional control (Montenegro-Montero & Larrondo 2016).
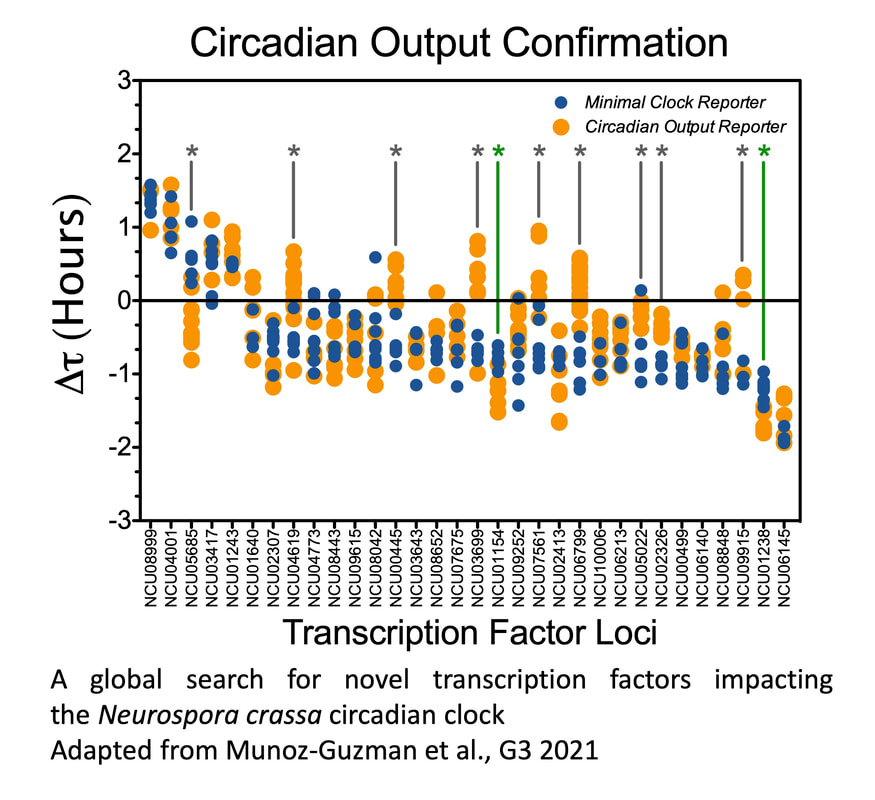
In an effort to identify additional transcriptional regulators modulating the N. crassa clock, we conducted a reverse genetic screen successfully covering close to 60% of the N. crassa Transcription Factors (TFs). Besides the canonical core clock components WC-1 and -2, none of the tested transcriptional regulators proved to be essential for rhythmicity. Nevertheless, we identified a set of 23 TFs that when absent lead to discrete, but significant, changes in circadian period. While the current level of analysis does not provide mechanistic information about how these new players modulate circadian parameters, the results of this screen reveal that an important number of light and clock-regulated TFs, involved in a plethora of processes, are capable of modulating the clockworks, further blurring the line between core-clock and input-output pathways (Munoz-Guzman et al. 2021). |
|
II. Semi-Synthetic Clocks: Transcriptional rewiring and circadian oscillators circuitry
To test whether the evolutionary conserved TTFL architecture is the only possible way to attain a functional eukaryotic circadian clock, we have adopted a transcriptional rewiring approach, forcefully co-opting regulators of the circadian output pathways into the core-oscillator. Our results show that one of the obtained semi-synthetic clocks is capable of maintaining all defining circadian properties yet, notably, it also exhibits new characteristics, providing new insights into the evolution of clock designs (Goity et al., submited). |
We are building other clock circuits combining synthetic promoters and defined cis-modules, obtaining semi-synthetic oscillators with notable changes in amplitude which can be modulated by external cues. Such results provide further proof of the genetic and transcriptional plasticity of circadian circuits and their ability to integrate complex transcriptional programs (Del Rio-Pinilla et al., in preparation).
Our general interest in gene regulation in Neurospora have additionally led us to be a part of a major effort aimed at understanding overall transcription factor specificity in eukaryotes (Weirauch et al., 2014).
Our general interest in gene regulation in Neurospora have additionally led us to be a part of a major effort aimed at understanding overall transcription factor specificity in eukaryotes (Weirauch et al., 2014).
|
III. Fungal Clocks, Light and Virulence
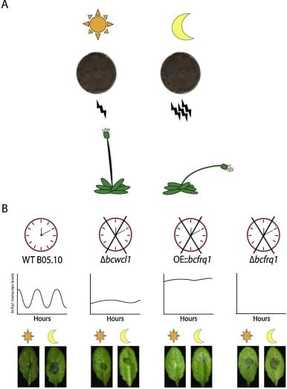
The circadian clock of the model plant Arabidopsis thaliana modulates defense mechanisms, which impact plant-pathogen interaction dynamics. The effect of the time of day on the virulence of plant pathogenic fungi on the other hand, has not been explored in detail and moreover, molecular description of clocks in fungi other than Neurospora have been largely neglected (Larrondo & Canessa 2019). We therefore focused on characterizing the circadian system of the plant pathogen Botrytis cinerea, one of the most scientifically/economically relevant plant pathogenic fungi. We have shown the existence of a functional clock in B. cinerea, which shares similar components and circuitry with the Neurospora circadian system (Hevia et al., 2015). Notably, we additionally described how circadian regulation affects the pathogenic potential in this fungus (Hevia et al., 2015; Hevia et al., 2016), such that the time of the day at which the first interaction between Botrytis and a plant (A. thaliana) occurs, defines the outcome of the interaction: bigger lesions are observed when the fungus interacts at dusk, compared to dawn. |
From Hevia et al. (2015). Circadian clocks and the regulation of virulence in fungi: Getting up to speed. Semin Cell Dev Biol. 57:147-55.
|
Unexpectedly, our Botrytis data show novel, extra-circadian functions for the conserved clock protein FRQ, something that we are actively pursuing (Muller-Esparza H, B. Sc. thesis 2015; Hevia et al., 2016). Indeed, our current studies have revealed a potential role for FRQ integrating signals of metabolic (nitrogen) and stress pathways (Seguel et al., in preparation).
In addition, as light is a key environmental cue that allows clock entrainment, we have been interested in different aspects of fungal photobiology, reporting for instance that light perception can modulate B. cinerea virulence (Canessa et al., 2013), while also analyzing the gene expression patterns affected by light, as this pathogen grows on plant tissue (Perez-Lara et al., 2023), .
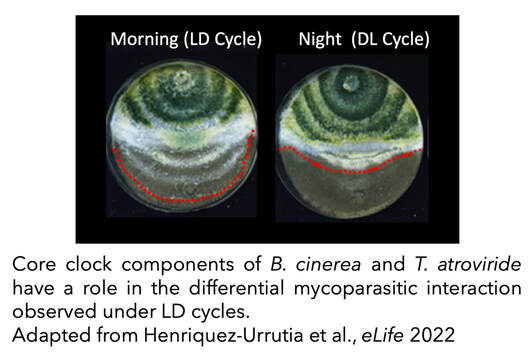
Moreover, the relevance of circadian clocks (or light) in fungal–fungal interactions has been also largely unexplored and, therefore, we sought to characterize a functional clock in the biocontrol agent Trichoderma atroviride to assess its importance in the mycoparasitic interaction against the phytopathogen B. cinerea. We confirmed the existence of circadian rhythms in T. atroviride, which are temperature-compensated and modulated by environmental cues such as light and temperature. Nevertheless, the presence of such molecular rhythms appears to be highly dependent on the nutritional composition of the media. Confrontation assays between wild-type and clock mutant strains of T. atroviride and B. cinerea, in constant light or darkness, revealed an inhibitory effect of light on T. atroviride’s mycoparasitic capabilities. Interestingly, when confrontation assays were performed under light/dark cycles, T. atroviride’s overgrowth capacity was enhanced when inoculations were at dawn compared to dusk. In addition, T. atroviride clock components largely modulate development and secondary metabolism in this fungus, including the rhythmic production of distinct volatile organic compounds (VOCs) (Henriquez-Urrutia et al., 2022).
In addition, as light is a key environmental cue that allows clock entrainment, we have been interested in different aspects of fungal photobiology, reporting for instance that light perception can modulate B. cinerea virulence (Canessa et al., 2013), while also analyzing the gene expression patterns affected by light, as this pathogen grows on plant tissue (Perez-Lara et al., 2023), .
Moreover, the relevance of circadian clocks (or light) in fungal–fungal interactions has been also largely unexplored and, therefore, we sought to characterize a functional clock in the biocontrol agent Trichoderma atroviride to assess its importance in the mycoparasitic interaction against the phytopathogen B. cinerea. We confirmed the existence of circadian rhythms in T. atroviride, which are temperature-compensated and modulated by environmental cues such as light and temperature. Nevertheless, the presence of such molecular rhythms appears to be highly dependent on the nutritional composition of the media. Confrontation assays between wild-type and clock mutant strains of T. atroviride and B. cinerea, in constant light or darkness, revealed an inhibitory effect of light on T. atroviride’s mycoparasitic capabilities. Interestingly, when confrontation assays were performed under light/dark cycles, T. atroviride’s overgrowth capacity was enhanced when inoculations were at dawn compared to dusk. In addition, T. atroviride clock components largely modulate development and secondary metabolism in this fungus, including the rhythmic production of distinct volatile organic compounds (VOCs) (Henriquez-Urrutia et al., 2022).
IV. Plant Cell Wall Deconstruction
We have participated in various fungal genome projects and we have been actively involved in the genome annotation of several fungi capable of lignin and cellulose degradation (Fernandez-Fueyo et al., 2012; Floudas et al., 2012; Hori et al., 2014)
In addition to these computational efforts, we have studied specific transcription factors (TFs) and environmental conditions modulating some of these processes, for instance, a URSB1/SRE-1 type TF and iron availability (Canessa et al., 2012; Canessa and Larrondo, 2013).
We have participated in various fungal genome projects and we have been actively involved in the genome annotation of several fungi capable of lignin and cellulose degradation (Fernandez-Fueyo et al., 2012; Floudas et al., 2012; Hori et al., 2014)
In addition to these computational efforts, we have studied specific transcription factors (TFs) and environmental conditions modulating some of these processes, for instance, a URSB1/SRE-1 type TF and iron availability (Canessa et al., 2012; Canessa and Larrondo, 2013).
We have additionally described that in N. crassa, the bZIP TF HAC-1 is involved in the unfolded protein response, and that this protein, and the ability to properly activate UPR, is key in allowing growth on media containing cellulose as the sole carbon source, unveiling new aspects of the degradation of this relevant plant cell-wall component (Montenegro-Montero et al., 2015).
We have devoted efforts to assess the effect of clock-regulation on the ability of Neurospora to degrade cellulose, confirming that the clock runs robustly as Neurospora grows on natural substrates such as plant material (Diaz & Larrondo 2020).
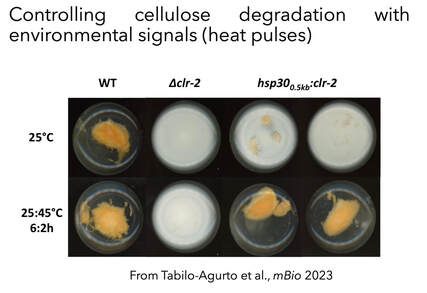
We have also developed new cis-modules capable of integrating environmental stimuli (i.e. temperature) to control, at will, cell wall deconstruction capabilities, as part of synthetic biology approach (Tabilo-Agurto et al., 2023). Such strains are being analyzed relative to their cellulolytic potential and their ability to degrade cellulose at higher temperatures.
We have devoted efforts to assess the effect of clock-regulation on the ability of Neurospora to degrade cellulose, confirming that the clock runs robustly as Neurospora grows on natural substrates such as plant material (Diaz & Larrondo 2020).
We have also developed new cis-modules capable of integrating environmental stimuli (i.e. temperature) to control, at will, cell wall deconstruction capabilities, as part of synthetic biology approach (Tabilo-Agurto et al., 2023). Such strains are being analyzed relative to their cellulolytic potential and their ability to degrade cellulose at higher temperatures.
|
V. Optogenetics and Synthetic Biology
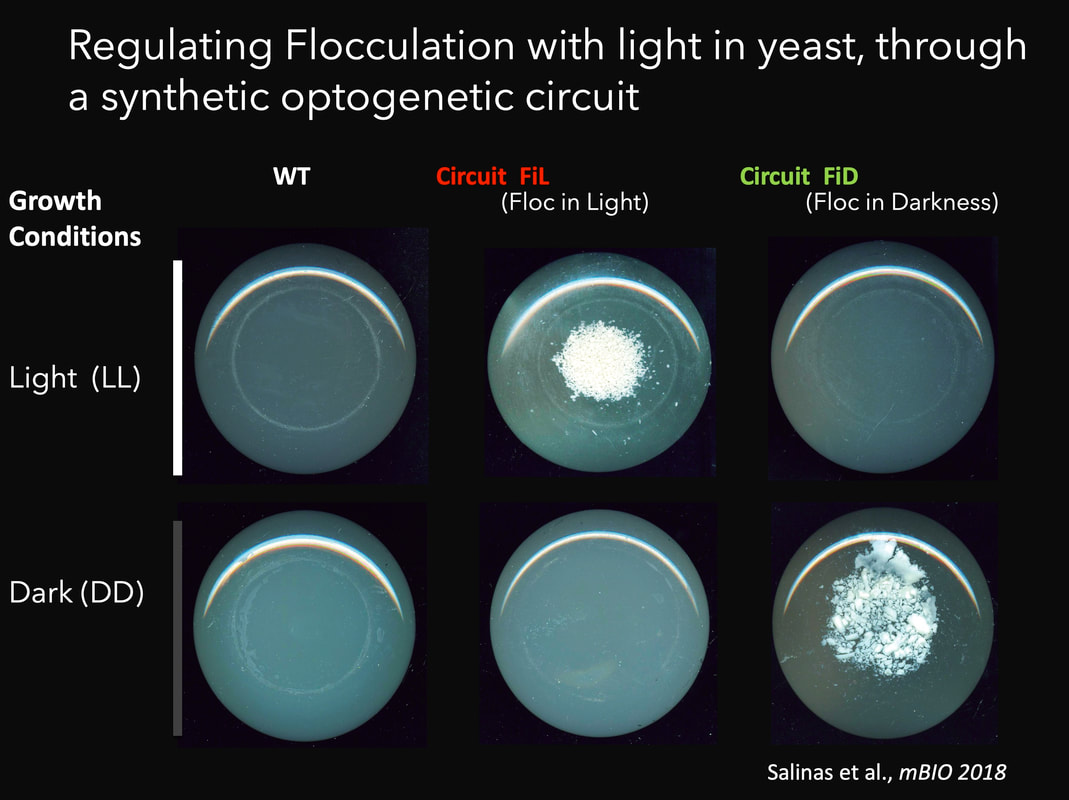
Together with synbio approaches aimed at modifying the Neurospora circadian system, we are also interested in designing and implementing additional circuits in both Neurospora and in S. cerevisiae. The interest in transcriptional processes mediated by light have led us to the development of optogenetics: the use of light to control cellular processes at will (Salinas et al., 2017). Thus, we have contributed to the design of transcriptional optogenetic switches (Romero et al., 2021; Rojas et al., 2022), which have high biotechnological potential , but at the same time allow for intricate experimental designs. Indeed, optogenetic switches are molecular devices which allow the control of different cellular processes by light, including gene expression, providing a poweful alternative to chemical inducers. We have developed novel optogenetic switches such as FUN-LOV, based on the LOV domain interaction of two blue-light photoreceptors (WC-1 and VVD) from N. crassa (Salinas et al., 2018; Romero et al., 2021). In yeast FUN-LOV provides tight regulation of gene expression, with low background levels in darkness and a highly dynamic and potent induction by light. We have used FUN-LOV to optogenetically manipulate in yeast biotechnologically relevant phenotypes, such as heterologous protein expression and flocculation, resulting in strains with potential industrial applications (Salinas et al., 2018). Cell communication is a widespread mechanism in biology, allowing the transmission of information about environmental conditions. We have coupled cell communication and optogenetics in S. cerevisiae, based on the light-dependent production of α-factor pheromone in one cell type, allowing the induction of gene expression in the other type. The results showed that external cues (light) is propagated through a diffusible signaling molecule to modulate gene expression in a synthetic system involving microbial cells, paving the road for studies allowing optogenetic control of population-level dynamics (Rojas et al., 2023). We are now interested in the implementation, in yeast, of more complex intercellular genetic circuits, for the analysis of population dynamics and synthetic microbial communities. In addition, some of our research, such as the “Live Canvas” project, lays at the interphase between fundamental science and art. Stay tuned to learn more about this and other ongoing projects. |